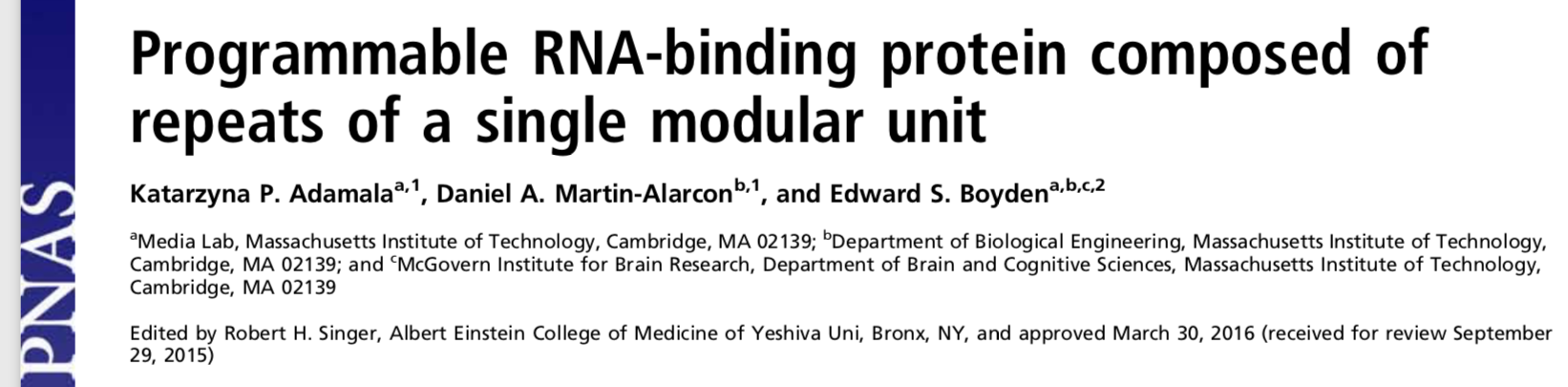
Pumby is a programmable protein designed to bind to the secondary genetic messenger, Ribonucleic Acid or RNA, in living mammalian cells. I designed and built Pumby for my PhD thesis while I worked in Ed Boyden’s group at the MIT Media Lab. You can always read the full published paper (PNAS copy; local copy), but I believe that every good paper deserves an informal recap of what the project was about and why it is interesting. That is what you’ll find here.
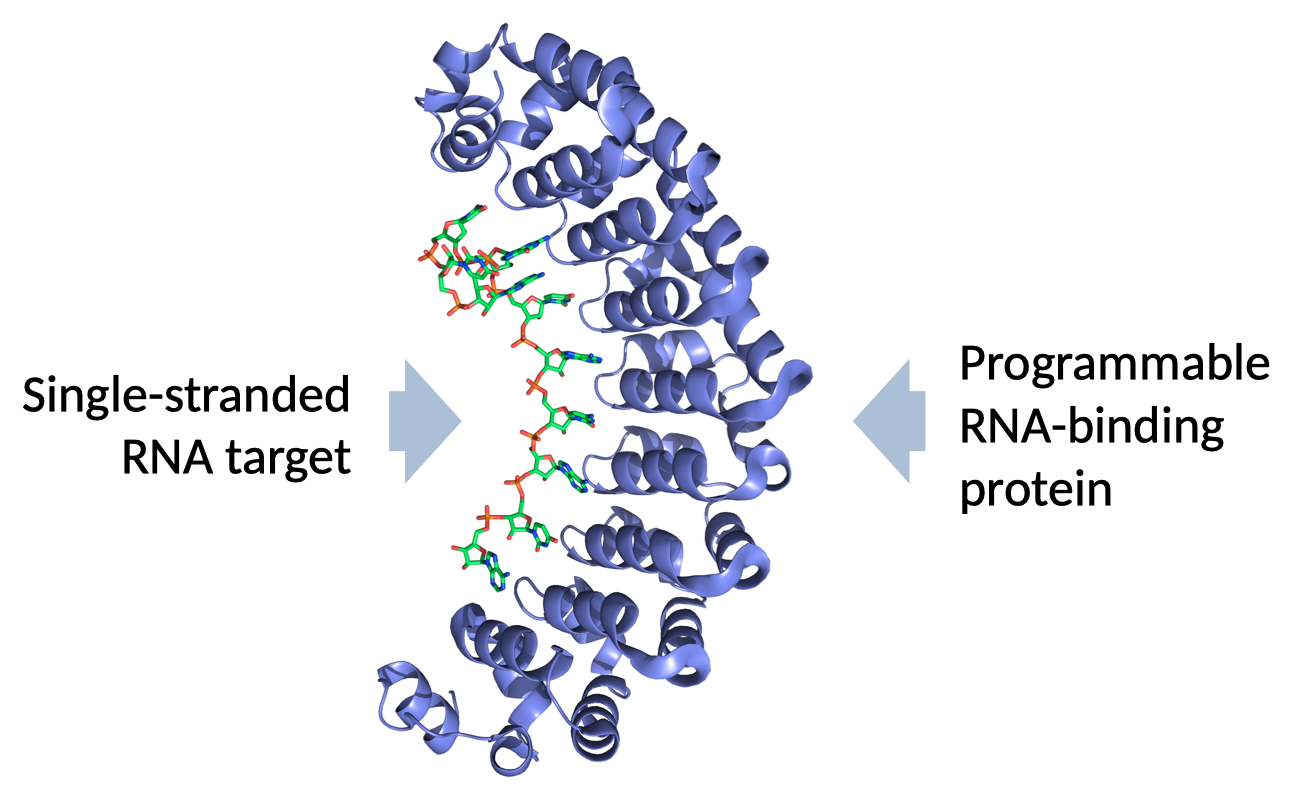
What is a programmable molecule?
Much of modern bioengineering is based on tools that bind to important biomolecules and execute some useful behavior, preferably in living organisms. The most famous biotechnology of the last decade, CRISPR, is based on a protein-RNA complex that can bind to specific sequences of DNA in the genome of basically anything. Many human diseases are caused by small errors in the genome. A protein that can specifically bind to those problematic sequences can also be used to edit them, thereby correcting the problem and eventually leading to a cure.
One of the great advantages of CRISPR is that it is a programmable complex. Many drugs work by binding to a very specific molecular target somewhere in the body. If the target changes or you want to treat a different disease, you have to design a completely different drug molecule. Programmable drugs, on the other hand, are those where the same molecule can be easily redesigned to bind to completely different targets. We created Pumby because we wanted a programmable molecule that could bind to RNA.
The dream: to use RNA as a molecular scaffold
Pumby is a general-purpose tool for synthetic biology, but we originally designed it to solve a very specific problem in neuroscience (the Boyden lab is, after all, focused on neurotech). Neurons are very dynamic systems, with lots of internal variables that change over time and affect each other. Neuroscientists frequently monitor these changes using molecules that change their fluorescence in response to some important variable. A calcium sensor like GCaMP shines bright when calcium levels spike, as happens in neurons when they talk to each other. In brains containing GCaMP, then, you can literally see brain activity in a microscope.
Fluorescent sensors like GCaMP are great, but they have a common problem: the best ones are all the same color. Two different sensors can report simultaneously, but if they’re evenly distributed throughout the cell then their signals will overlap and be very difficult to tell apart. I prefer to express this idea in haiku:
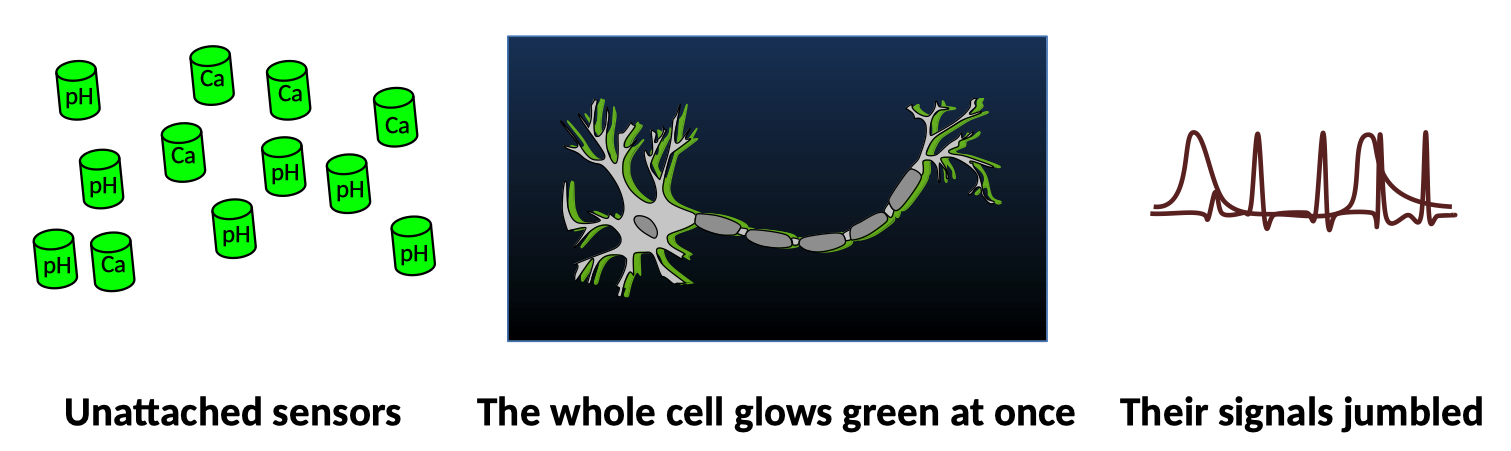
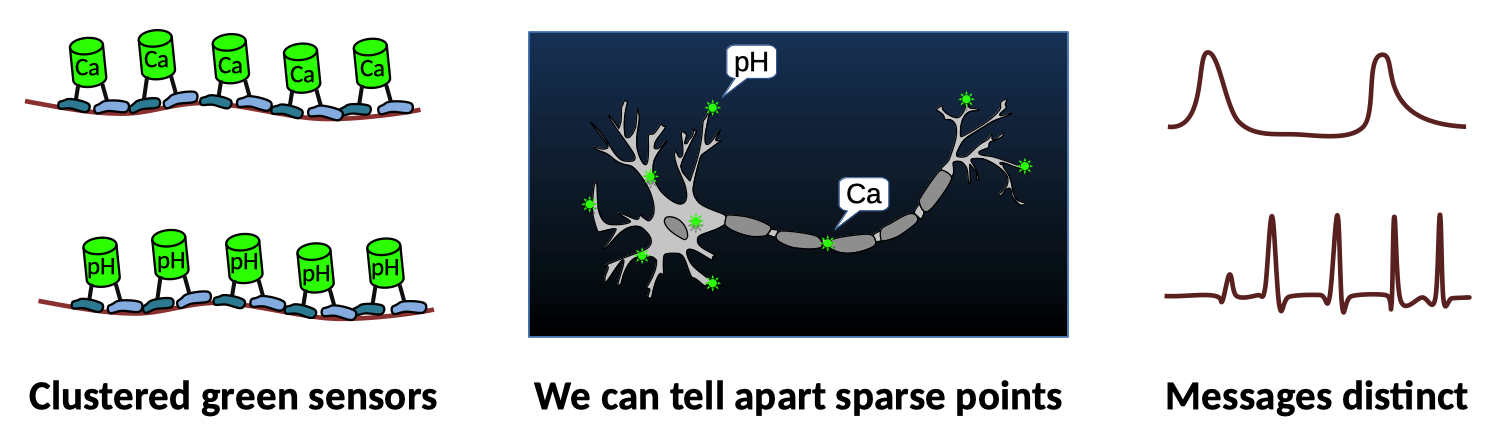
What if, however, sensors of a given type clustered together? If you attach sensors to Pumby, and express an RNA molecule with many copies of the sequence that Pumby recognizes, you could build an RNA scaffold that will help the sensors cluster together. Clustered sensors then have two distinct advantages. First, clusters are way brighter than the same number of sensors spread throughout the whole cell. Second, clusters of different sensor types are separate from each other and can be monitored independently with the same microscope. And so, Pumby expands the reach of some of the most widely used tools in neuroscience.
Building and testing a new protein
We started out by adapting a well-studied human protein called the Pumilio Homology Domain, or PumHD. This protein contains 8 units that each bind a particular RNA base, so that the overall protein binds a single 8-letter sequence of RNA (AUAGAUGU). PumHD also contains two non-binding caps, which we number 0 and 9. The 8 binding units are very similar, and they all interface with RNA through a specific set of 3 key amino acids, two that touch the base directly and one that stacks in between bases.
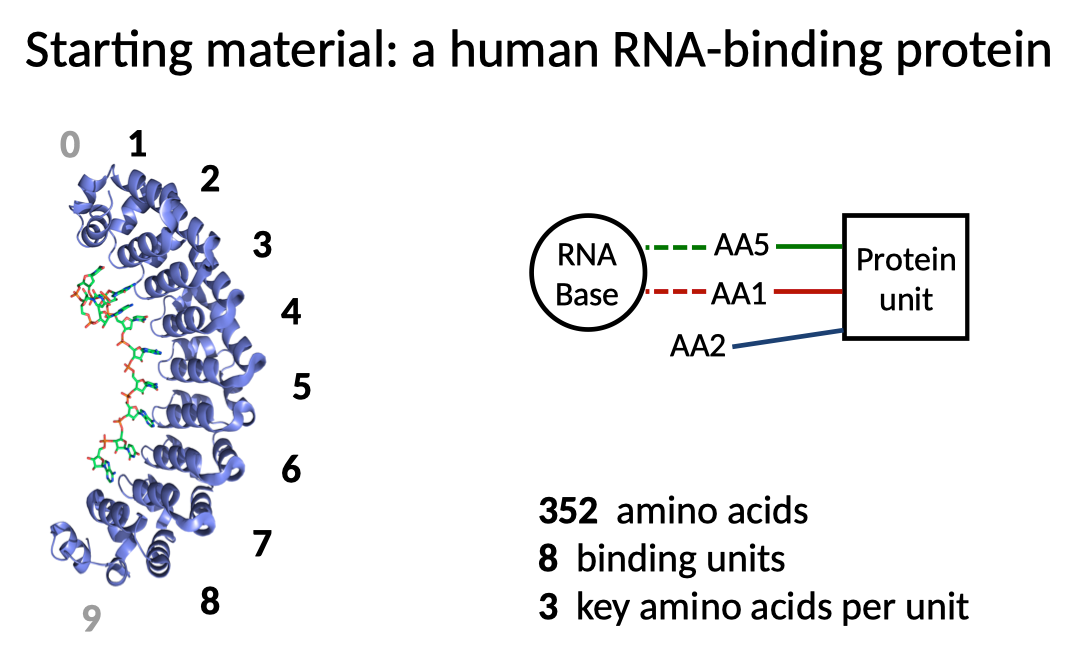
We first figured out how to make custom PumHDs with altered binding at each of the 8 units, so that each unit could be designed to bind each of the 4 RNA letters. We did this by testing many combinations of key amino acids, and determining which key amino acids would do the trick at that position. Our custom PumHD molecules can thus be designed to bind arbitrary 8-letter RNA sequences.
The real advancement, however, was to create a more versatile descendant of PumHD. We discovered that it’s possible to concatenate a single unit of PumHD into chains of arbitrary length. These chains proved to be stable despite the absence of cap units, and despite the fact that they didn’t have the diversity of PumHD with its 8 different units. We found that we could build stable chains with any length between 6 and 18 units. Each of the units can be chosen to bind any RNA base, so that the overall molecule can bind arbitrary RNA sequences between 6 and 18 letters long. We called our method Pumilio-Based Assembly, or Pumby.
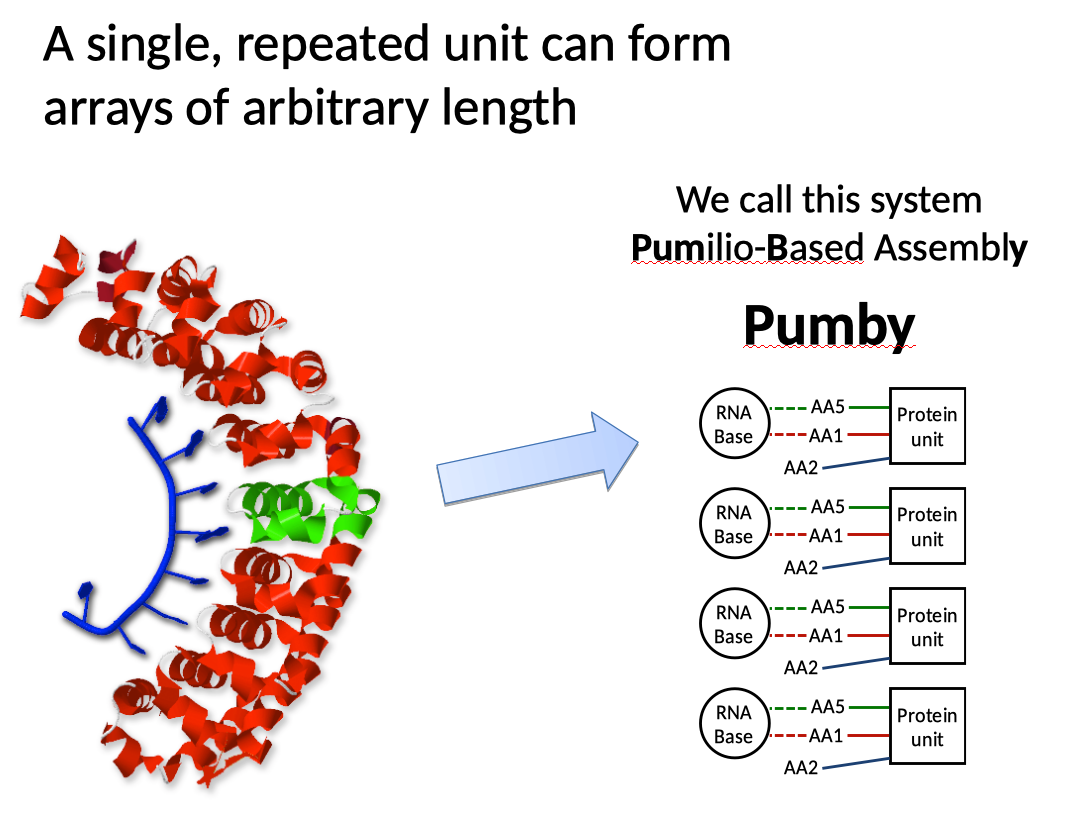
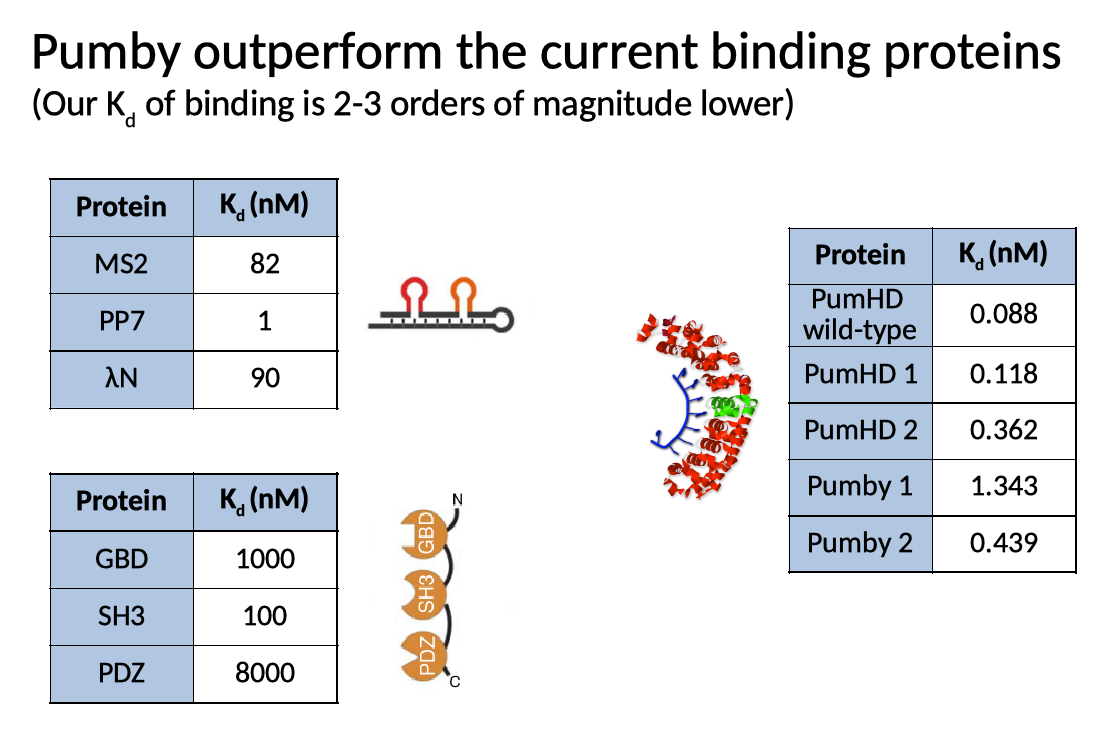
We put our molecules through a variety of tests in cultured mammalian cells. We showed that Pumby behaved equivalently to our custom PumHD molecules in a variety of settings, and that both bound their targets with an affinity equivalent to that of wild-type PumHD. More importantly, our proteins showed much greater binding affinity than other protein domains normally used in synthetic biology for similar applications.
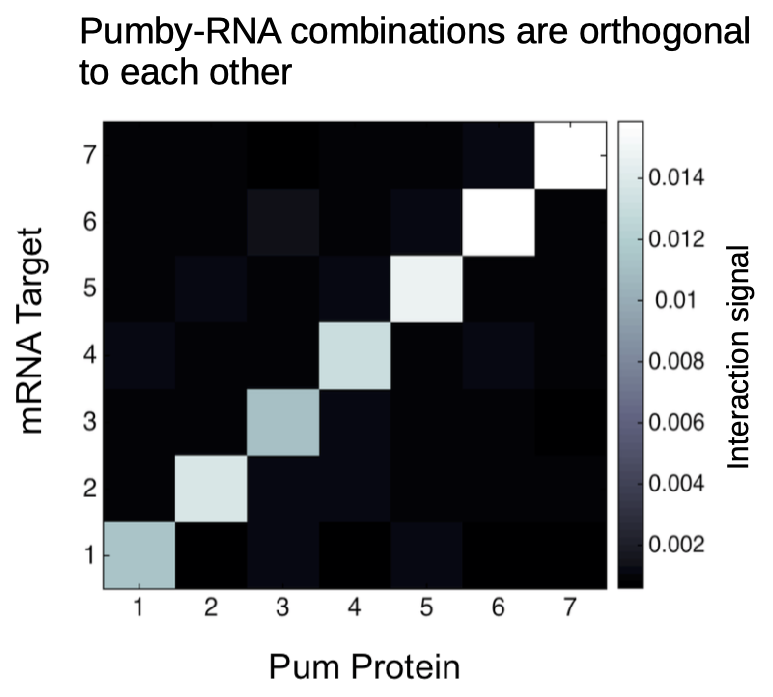
But even more important than binding affinity is orthogonality. Two Pumby proteins, once designed to match two different targets, can coexist in the same cell without interference. In fact, one can express several proteins in the same cell at the same time and have them each perform an independent function. We demonstrated this orthogonality with an assay where 7 Pums (some PumHD, some Pumby) were exposed to 7 transcripts, under conditions where binding to the target would produce a signal.
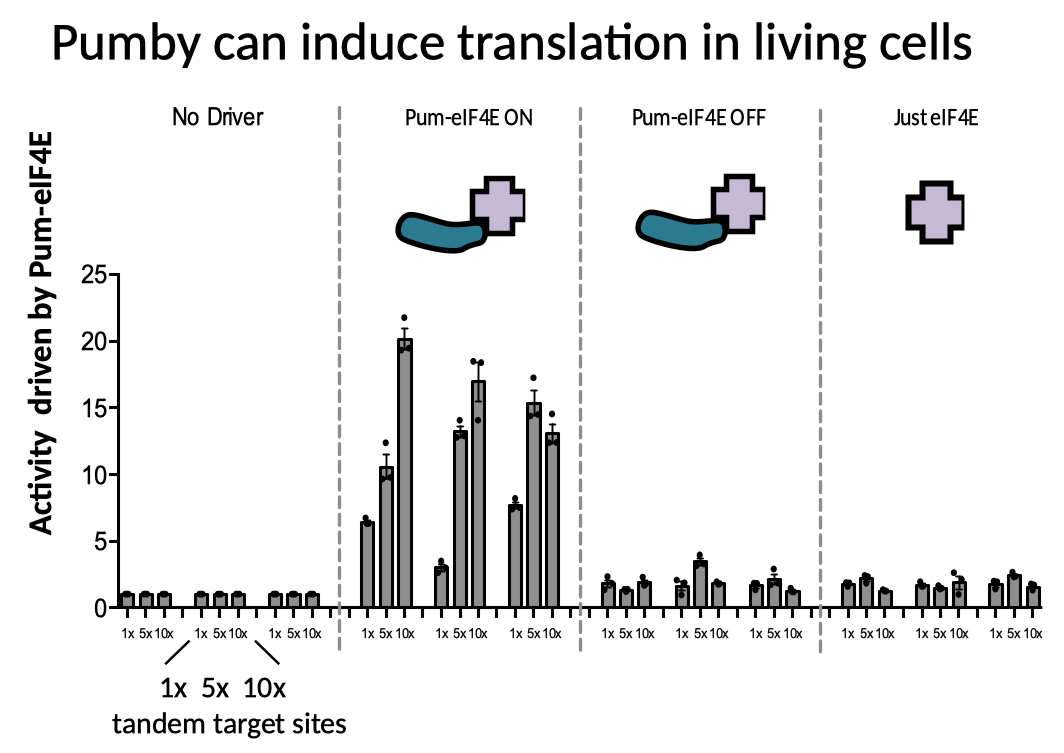
In separate experiments detailed in our paper, we showed that Pumby proteins can be used to detect the presence of particular transcripts in cells, to measure the translation of those transcripts, and even to induce translation by recruiting a translation driver (eIF4E) to a location on a transcript just upstream of an open reading frame.
In conclusion
What we have created, then, is a method for specifically binding arbitrary sequences of RNA in living cells. One could use Pumby to interact with biologically meaningful transcripts in living cells, altering gene expression at the transcriptional level without having to modify the genome itself. Alternatively, one could use synthetic RNA as I mentioned earlier, as a scaffold that can be combined with Pumby proteins to create programmable sub-cellular structures. As is always the case in synthetic biology, we’re excited to see which applications will prove to be the most useful.
